1 Overview
XpressWeed is a texture-prior driven segmentation framework for weed identification in precision agriculture. It reframes the problem: instead of treating plant leaves as rigid objects, XpressWeed models them as textures — leveraging the fact that each species exhibits distinctive vein patterns, surface roughness, and micro-structures that remain stable under deformation, occlusion, and changing illumination.
A Fully Convolutional Network (FCN) is pre-trained on collages from 12 leaf texture classes, building robust and species-agnostic feature representations. A Model-Agnostic Meta-Learning (MAML) strategy then adapts this model to 4 completely unseen weed species using only 80 images — achieving 85% accuracy versus 48% for traditional MAML on the same data.
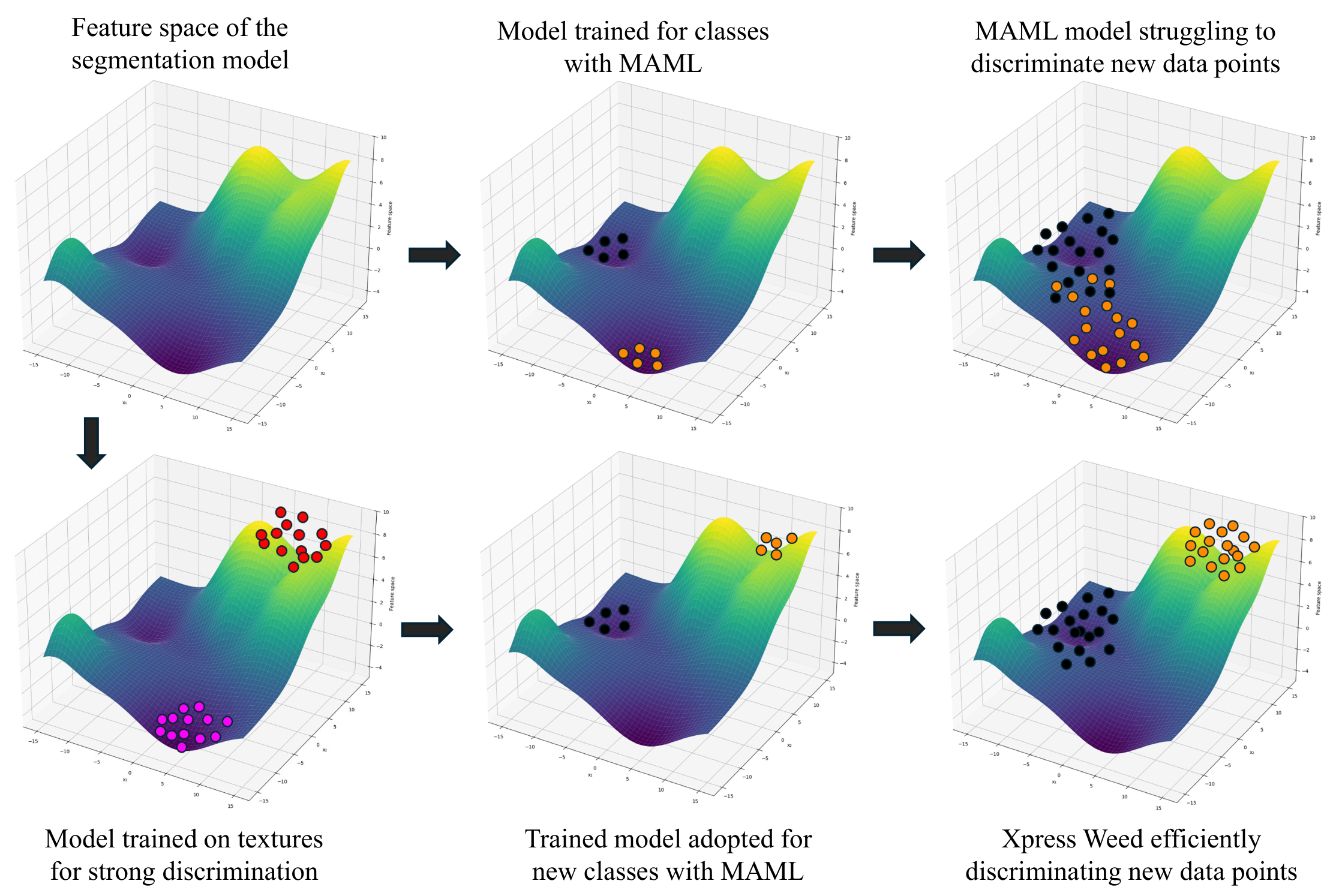
2 The Problem
Weeds compete with primary crops for nutrients, water, and sunlight — causing significant yield loss worldwide. While deep learning has been applied to weed segmentation, two fundamental obstacles remain unsolved:
Challenge: Scale
- Weed species vary across regions and seasons
- One large model for all species is impractical
- Requires exhaustive labeled datasets
- Computationally infeasible to retrain per region
Challenge: Variation
- Leaves overlap, twist, and change orientation
- Lighting shifts color hues dramatically
- Object-centric CNNs rely on shape — which fails
- Few-shot methods can't capture this variation
XpressWeed: Texture Priors
- Leaf textures are stable under deformations
- Vein patterns invariant to shape changes
- Pre-training on textures → species-agnostic filters
- No shape-based assumptions needed
XpressWeed: MAML Adaptation
- Texture priors = strong initialization for MAML
- Adapt to new species in 80 images
- N-way episodic training = implicit contrastive learning
- No expensive full-pipeline meta-training
3 Proposed Method
XpressWeed combines supervised texture pre-training with MAML-based few-shot adaptation. The two-stage design separates large-scale representation learning from lightweight few-shot adaptation, achieving both robustness and computational efficiency.
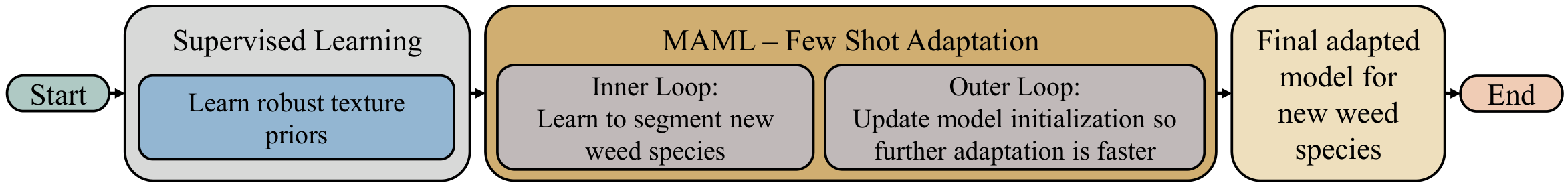
Pipeline Overview
12 species → Texture Collages
Overlap simulation → FCN Pre-training
Supervised + WCE → Texture Priors
Robust initialization → MAML Inner Loop
Support set update → MAML Outer Loop
Query meta-update → Adapted Model
4 unseen weeds
Stage 1 — Supervised Texture Pre-training
The FCN is trained on texture-prior images constructed from PlantVillage leaf patches arranged into collages. Overlapping leaves and background-removed variants simulate real farmland conditions where boundaries are difficult to separate. Weighted cross-entropy loss handles severe class imbalance. Meta-learning is deliberately not used at this stage — it would require expensive second-order differentiation over thousands of images.
Stage 2 — MAML-Based Adaptation
After pre-training, MAML adapts the model to 4 unseen weed species with only 80 labeled patches. Each episode uses 3-way classification, 12 support samples per class (inner loop update), and 10 query samples per class (outer loop meta-update). The n-way episodic setup implicitly mimics contrastive learning — same-class embeddings cluster together while different classes are pushed apart — without explicit contrastive losses.
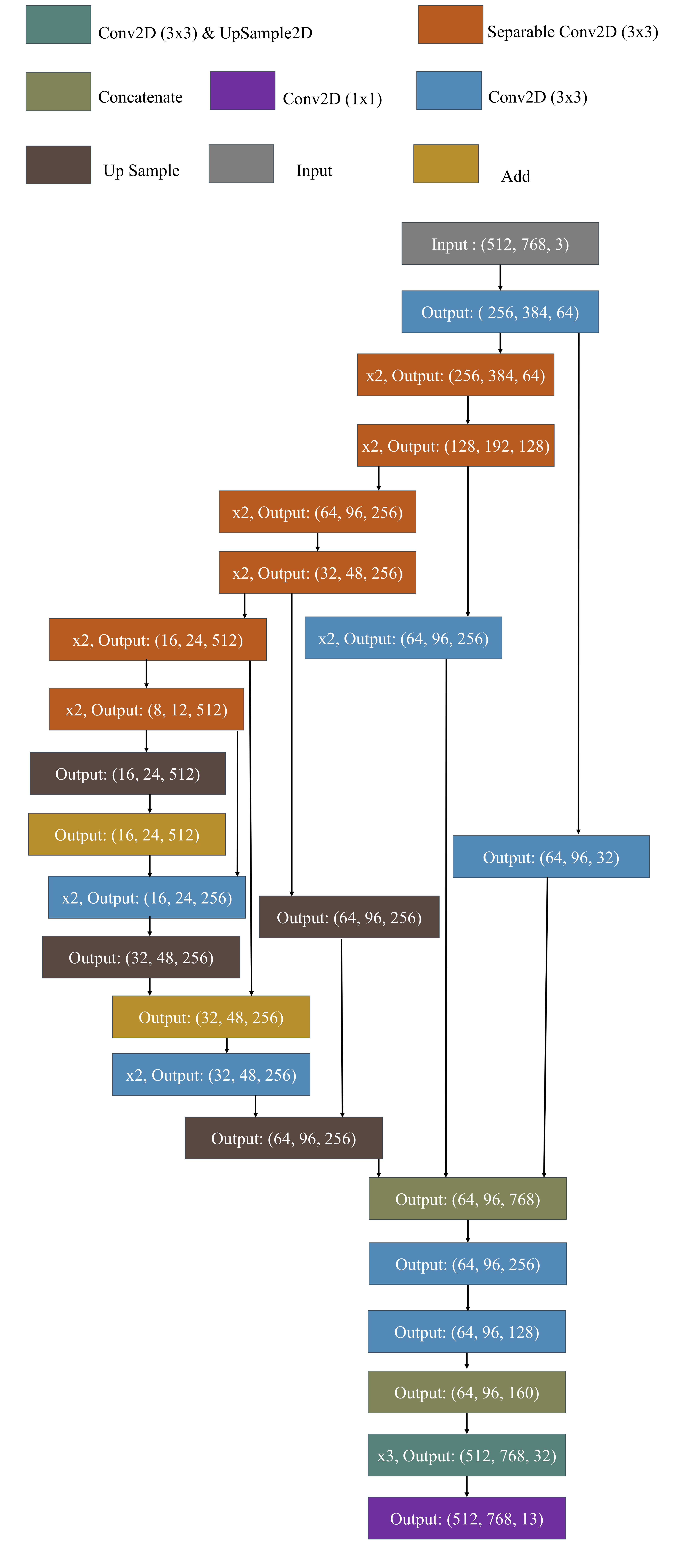
4 Network Architecture
The backbone is an encoder–decoder design inspired by U-Net, modified to emphasize texture discrimination and multi-scale feature fusion. It is intentionally lightweight — MAML's second-order gradient updates require smaller models to converge efficiently.
Encoder
Progressively downsamples input (512×768×3) through stacked conv blocks to a compact bottleneck (8×12×512). Captures high-level texture features stable under deformation and lighting changes. Skip connections preserve feature maps at each scale.
Decoder
Upsampled feature maps are concatenated with saved encoder maps at matching scales — multi-scale fusion provides both global texture context and fine-grained leaf-edge/venation details. Boundary refinement is achieved implicitly through feature fusion, not expensive deconvolution.
Output Head
A pointwise (1×1) convolution produces dense per-pixel segmentation masks, informed by features across all scales.
- Separable convolutions — reduce parameter count for faster MAML updates
- Skip connections — fuse global texture + fine-grained edges/venation
- Feature fusion — not transposed conv; simpler and effective
- Lightweight bottleneck — 8×12×512 compresses to essential texture features
- Weighted CE loss — handles severe weed/crop pixel imbalance
5 Stage 1: Pre-training Results
The FCN was trained on a PlantVillage-derived dataset of 12 leaf texture classes arranged into collages. The design intentionally introduces challenging conditions — overlapping leaves, background-removed variants, and mixed textures — to simulate real farmland boundaries.

Pre-training Metrics
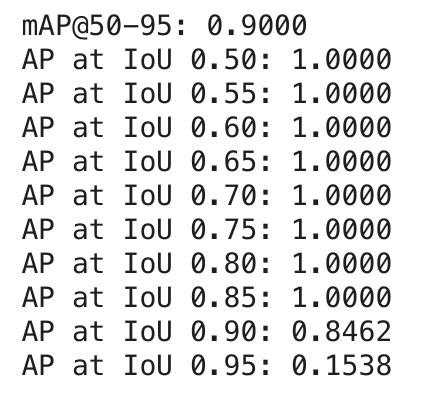
6 Stage 2: MAML Adaptation Results
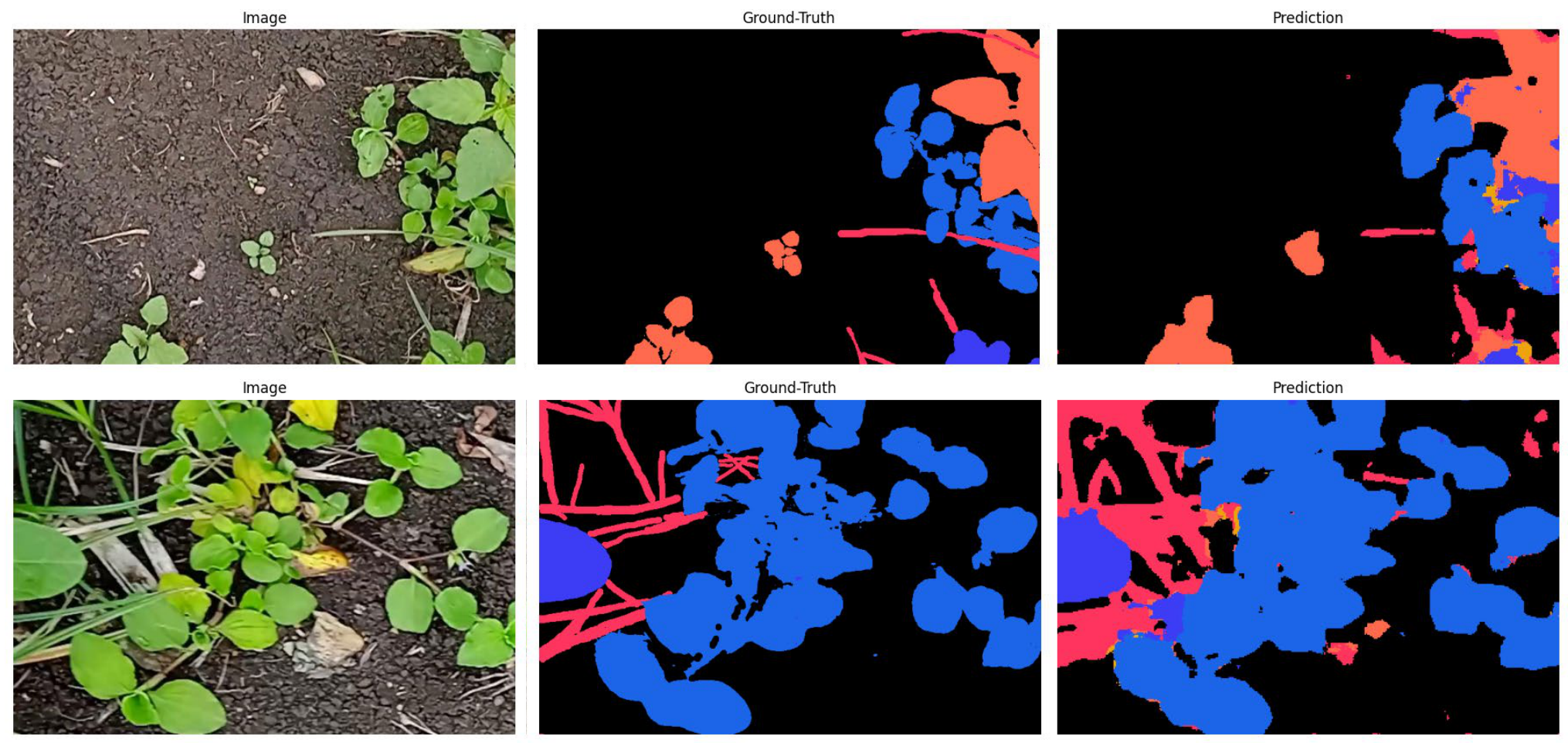
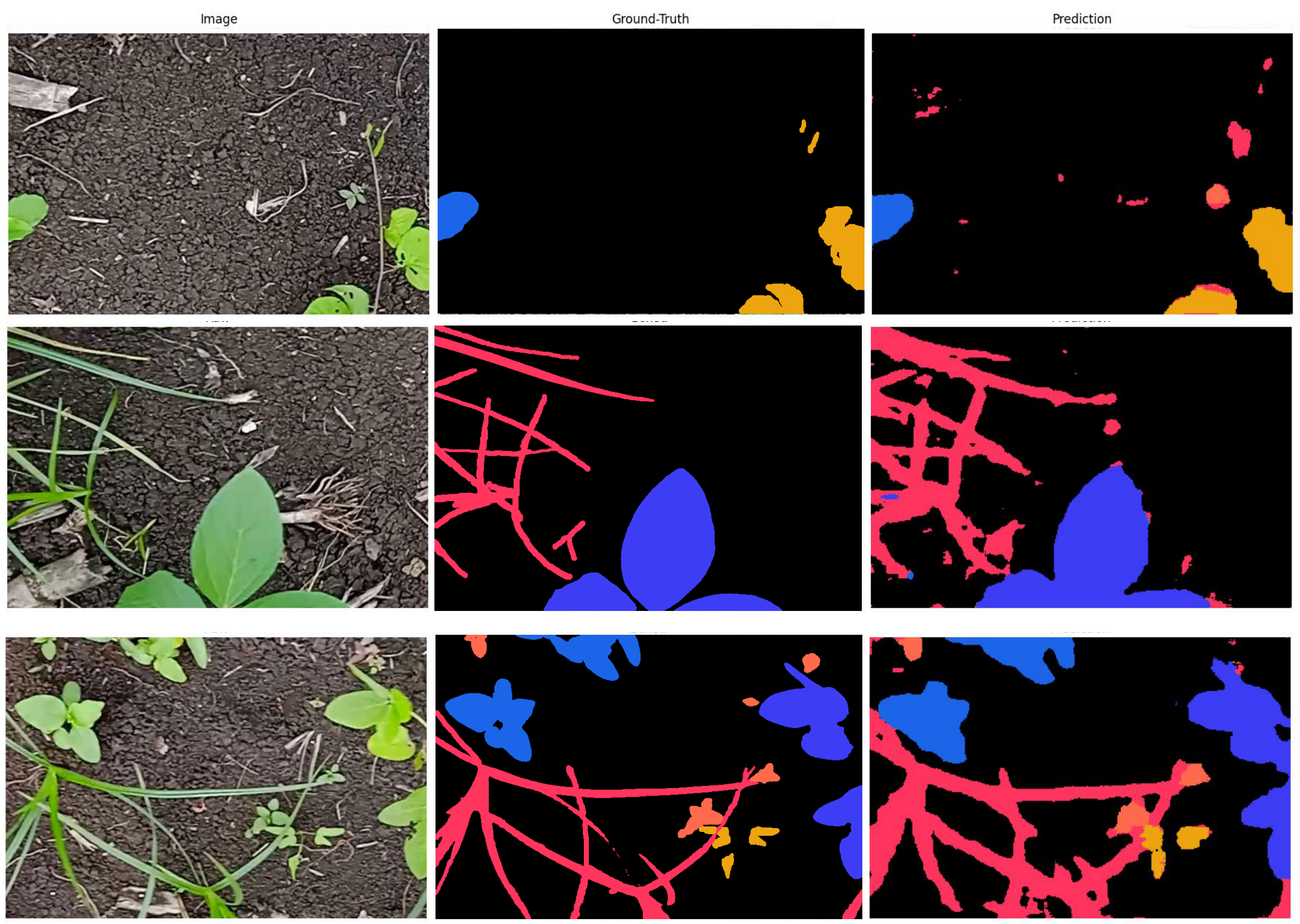
The pre-trained model is adapted to 4 unseen weed species using a real Indian farmland dataset. 20 high-resolution (2112×1600) images were cropped into 80 patches (512×768) for MAML training. The task is 3-way classification, 12-shot, over 25 epochs.

XpressWeed vs. Traditional MAML
Accuracy
Same network · Same 80 images · Different initialization
Mean IoU
Train set: 92% acc / 72% mIoU | Val set: 85% acc / 66% mIoU


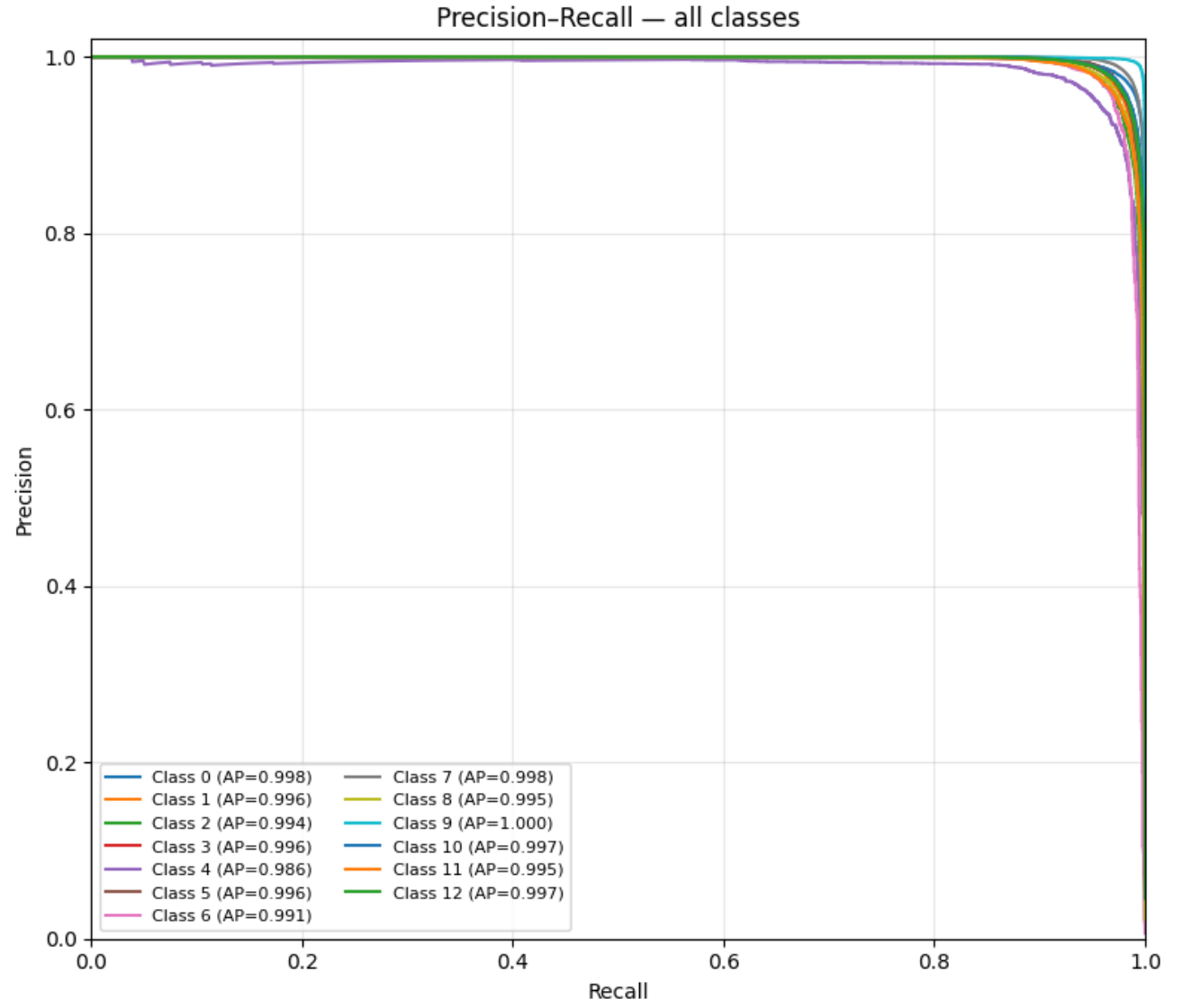
Per-Class Average Precision
| Summary Metric | Value |
|---|---|
| Macro mAP | 0.783 |
| Micro AUPRC | 0.962 |
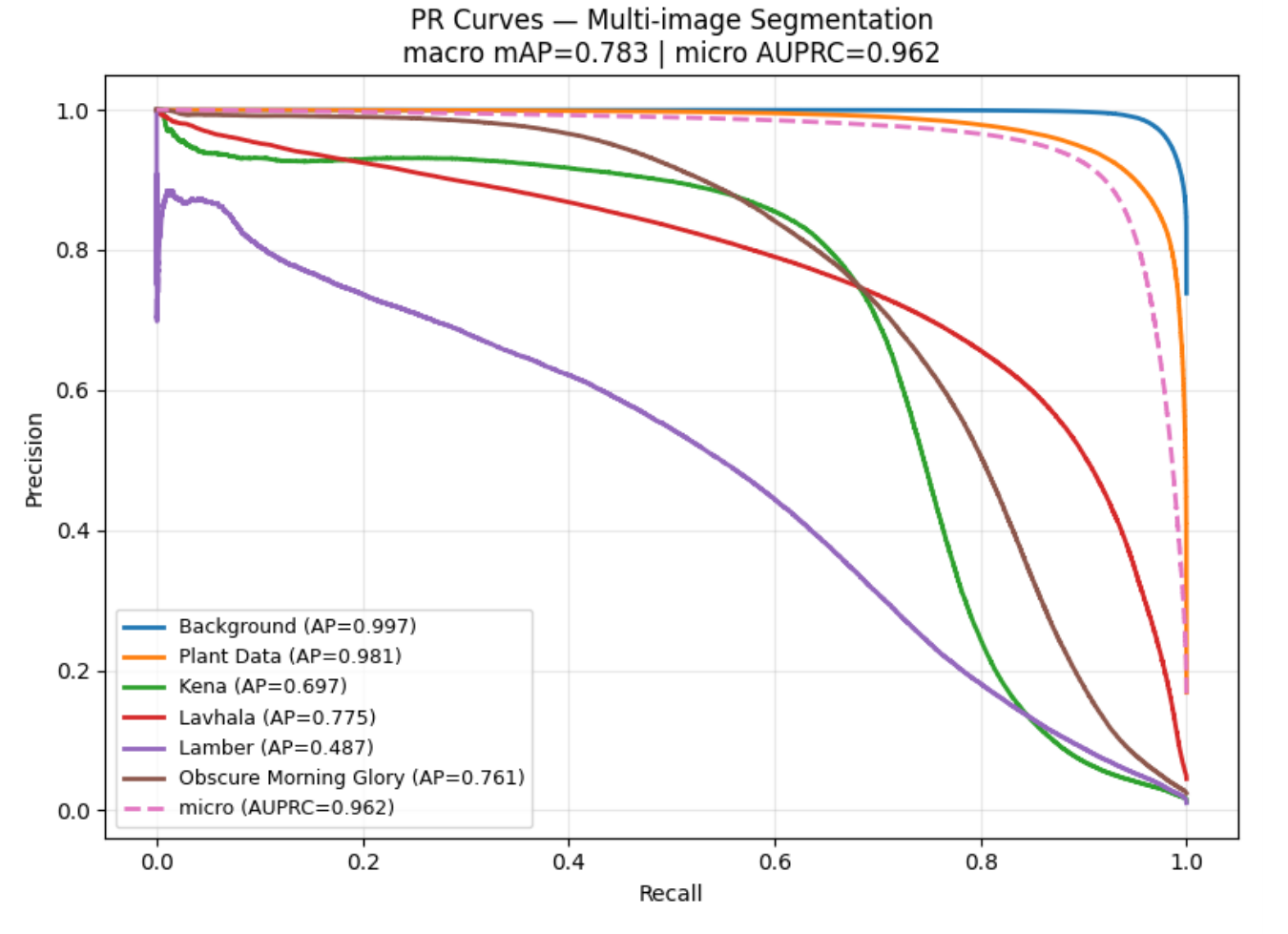
7 Comparison with Related Work
| Method | Year | Approach | Limitation vs XpressWeed |
|---|---|---|---|
| Syed & Suganthi | 2023 | Fuzzy active contour | No deep features; poor scalability |
| Naik / Hu et al. | 2024/22 | U-Net / U-Net++ | Static; requires large labeled datasets |
| Li et al. | 2023 | PSPNet | Static; cannot adapt to new classes |
| Shorewala et al. | 2021 | Semi-supervised ResNet-SVM | Binary classification only |
| Amac et al. | 2021 | Self-supervised MAML | Depends on saliency mask quality |
| Cao et al. | 2019 | FCN + Meta-Seg | Fails on plant image variability |
| Logeshwaran et al. | 2024 | Meta-stacking | Computationally heavy full-pipeline meta-learning |
| XpressWeed (Ours) | 2026 | Texture pre-training + MAML | 85% acc · 80 images · 4 unseen species · resource-efficient |
8 Code & Repository
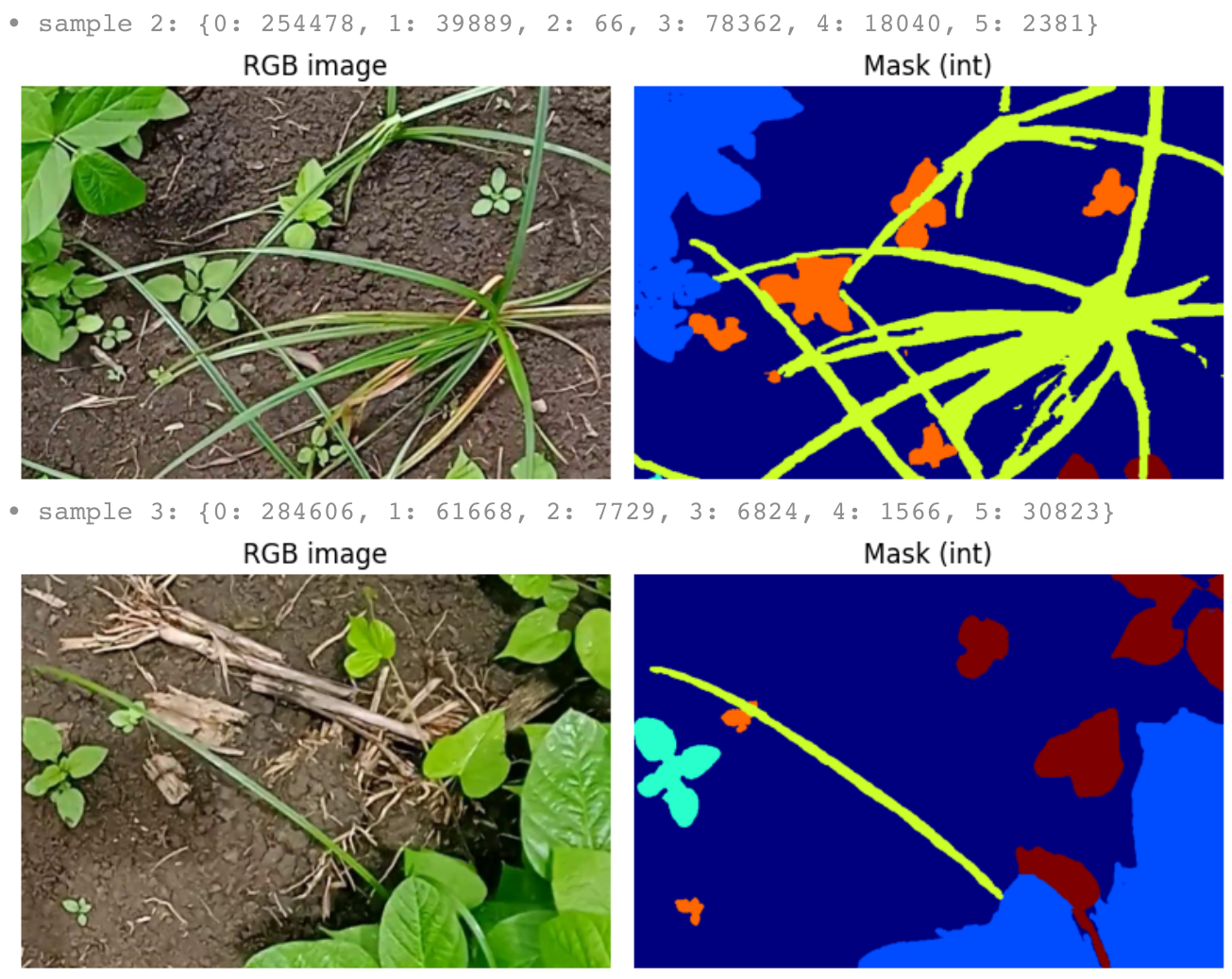
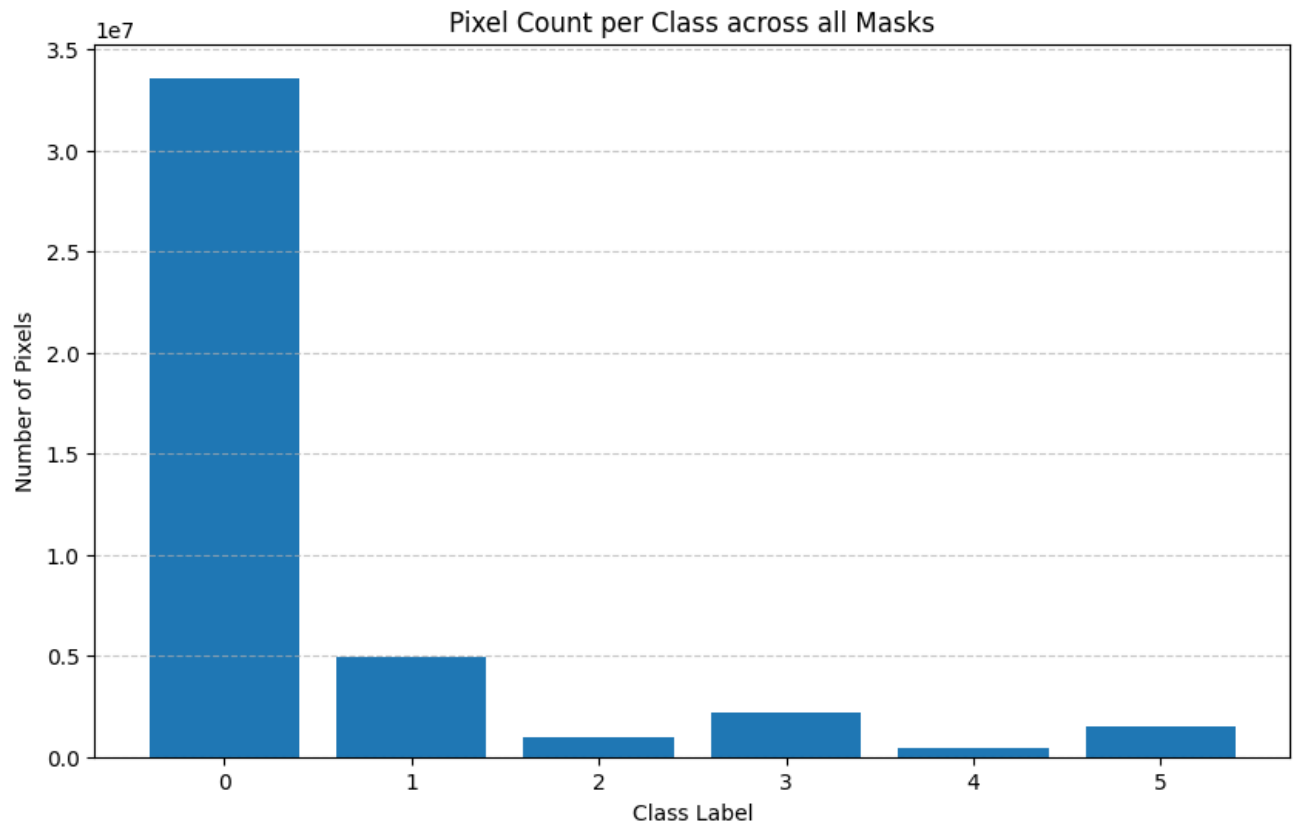
Class Color Mapping
# RGB mask colors → integer class IDs (farmland adaptation dataset) color2class = { (0, 0, 0): 0, # Background (61, 61, 245): 1, # Plant data (Soybean — seen in pre-training) (28, 101, 232): 2, # Kena (unseen weed) (255, 53, 94): 3, # Lavhala (unseen weed) (255, 106, 77): 4, # Lamber (unseen weed — fewest pixels) (238, 164, 15): 5, # Obscure Morning Glory (unseen weed) }
MAML Episode Structure
# Episode configuration N_WAY = 3 # classes per episode K_SUPPORT = 12 # support samples per class (inner loop) K_QUERY = 10 # query samples per class (outer loop) N_EPOCHS = 25 for epoch in range(N_EPOCHS): for episode in sample_episodes(): support_x, support_y = episode['support'] # (36, 512, 768, 3) query_x, query_y = episode['query'] # (30, 512, 768, 3) # Inner loop: fast adaptation on support set fast_weights = inner_loop_update(model, support_x, support_y) # Outer loop: meta-gradient update on query set meta_loss = compute_loss(fast_weights, query_x, query_y) meta_update(model, meta_loss)
Data Augmentation
# Augmentations applied consistently to both images AND masks: # 1. Random horizontal flip (p = 0.5) # 2. Random zoom 0.9× – 1.1× → crop-centre or reflect-pad # 3. Random shadow / highlight (RSH) (p = 0.85, triangular regions) # 4. Color jitter → hue, saturation, brightness # 5. Gaussian blur → simulates out-of-focus leaves # 6. Gaussian noise → simulates sensor noise aug_imgs, aug_masks = augment_images_and_masks( images_batch, masks_batch, target_size=(512, 768), rsh_prob=0.85 )
9 Citation
If you use XpressWeed in your research, please cite:
@inproceedings{kethineni2026xpressweed,
title = {XpressWeed: Meta-Inspired Few-Shot Adaptation for
Plant Weed Segmentation Using Texture Priors},
author = {Kethineni, Kiran K. and Kanukuntla, Rishi Raj and
Mohanty, Saraju P. and Kougianos, Elias},
booktitle = {Proceedings of the IEEE SusTech 2026},
year = {2026},
institution = {University of North Texas}
}